Today’s blog is about realistic communication, i.e., what kind of performance can realistically be expected of a wireless network. To get started, let’s have a look at an excerpt from a recent workshop description:
“Future wireless networks will have to support many innovative vertical services, each with its own specific requirements, e.g.
- End-to-end latency of 1 ns and reliability higher than 99.999% for URLLCs.
- Terminal densities of 1 million of terminals per square kilometer for massive IoT applications.
- Per-user data-rate of the order of Terabit/s for broadband applications.”
Let’s break this down, bullet by bullet.
First bullet: In 1 ns, light travels 30 cm in free space. So “end-to-end” here would mean a distance of at most 10 cm, to leave some fraction of a nanosecond for encoding, transmission, and decoding. But what useful wireless service is there where transceivers are within at most 10 cm? Next, a packet loss rate of 10-5 means that the spectral efficiency must be very low. Together with a latency constraint of 1 ns, ultrahigh bandwidths must be used, which, in turn, makes the design of circuitry and antenna arrays extremely challenging. At least the channel can be expected to be benign (line-of-sight).
Where does stochastic geometry come in? Assuming that these ultrashort links live in a network and not in isolation, interference will play a role. Let us consider a Poisson bipolar network with normalized link distance 1, a path loss exponent α and Rayleigh fading. What is the maximum density of links that can be supported that have an outage of at most ε? This quantity is known as the spatial outage capacity (SOC). For small ε, which is our regime of interest here, we have
where δ=2/α and cδ is a constant that only depends on the path loss exponent 2/δ. ρ is the spectral efficiency (in bits/s/Hz or bps/Hz). This shows the fundamental tradeoff between outage and spectral efficiency: Reducing the outage by a factor of 10 reduces the rate of transmission by the same factor if the same link density is to be maintained. Compared to a more standard outage constraint of 5%, this means that the rate must be reduced by a factor 5,000 to accommodate the 99.999% reliability requirement. Now, say we have 0.5 ns for the transmission of a message of 50 bits, the rate is 100 Gbps. Assuming a very generous spectral efficiency of 100 bps/Hz for a system operating at 5% outage, this means that 100 Gbps must be achieved at a spectral efficiency of a mere 0.02 bps/Hz. So we would need 5 THz of bandwidth to communicate a few dozen bits over 10 cm.
Even relaxing the latency constraint to 1 μs still requires 5 GHz of bandwidth.
In cellular networks, the outage-rate relationship is governed by a very similar tradeoff.
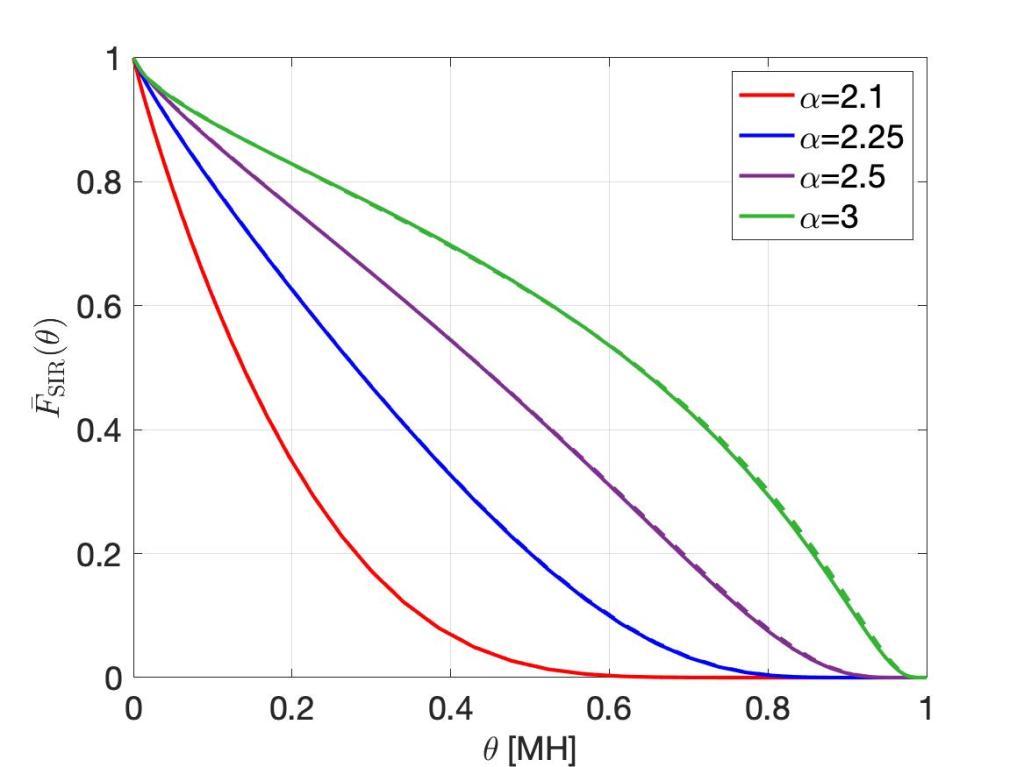
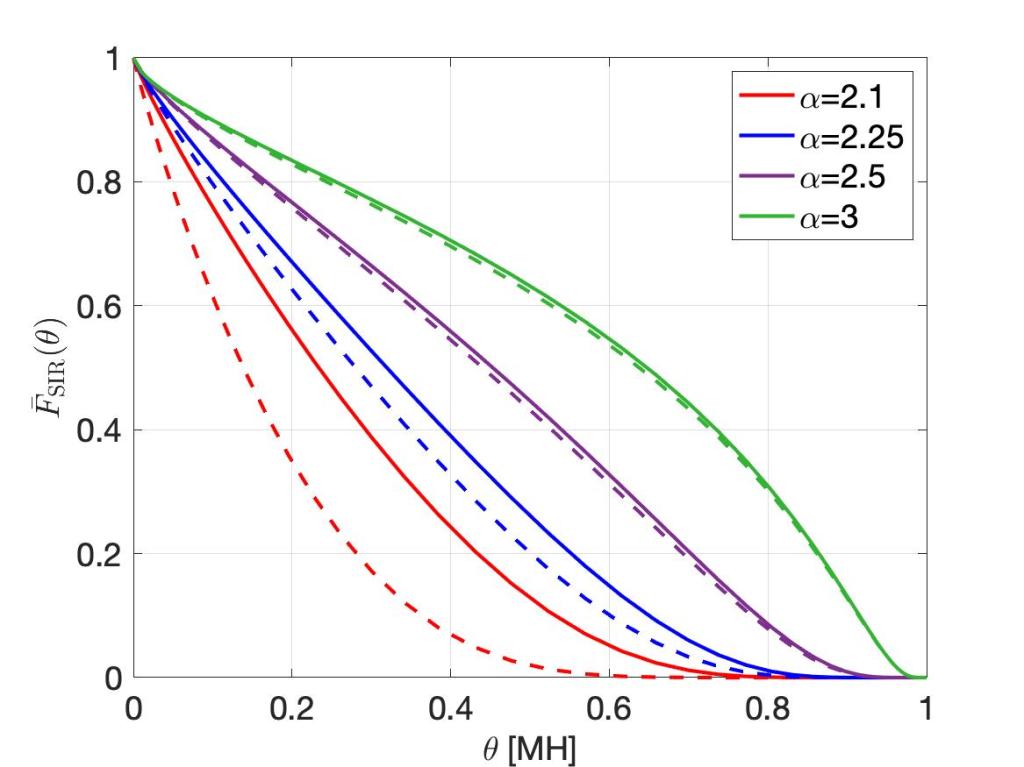
For any stationary point process of base stations and Rayleigh fading, the SIR meta distribution asymptotically has the form
where Cδ again depends only on the path loss exponent. This is the fraction of users who achieve a spectral efficiency of ρ with an outage less than ε, remarkably similar to the bipolar result. To keep this fraction fixed at, say, 95%, again the spectral efficiency needs to be reduced in proportion to a reduction of the outage constraint ε.
Second bullet: Per the classification and nomenclature in a dense debate, this density falls squarely in the tremendously dense class, above super-high density and extremely high density. So what do the anticipated 100 devices in an average home or 10,000 devices in an average parking lot do? What kind of messages are they exchanging or reporting to a hub? How often? What limits the performance? These devices are often said to be “connected“, without any specification what that means. Only once this is clarified, a discussion can ensue whether such tremendous densities are realistic.
Third bullet: Terabit-per-second (Tbps) rates require at least 10 GHz of spectrum, optimistically. 5G in its most ambitious configuration, ignoring interference, has a spectral efficiency of about 50 bps/Hz, and, barring any revolutionary breakthrough, more than 100 bps/Hz does not appear feasible in the next decade. Similarly, handling a signal 10 GHz wide would be an order of magnitude beyond what is currently possible. Plus such large junks of spectrum are not even available at 60 GHz (the current mm-wave bands). At 100 GHz and above, link distances are even more limited and more strongly subject to blockages, and analog beamforming circuitry becomes much more challenging and power-hungry. Most importantly, though, peak rates are hardly achieved in reality. In the 5G standard, the user experienced data rate (the rate of the 5-th percentile user) is a mere 1% of the peak rate, and this fraction has steadily decreased over the cellular generations:
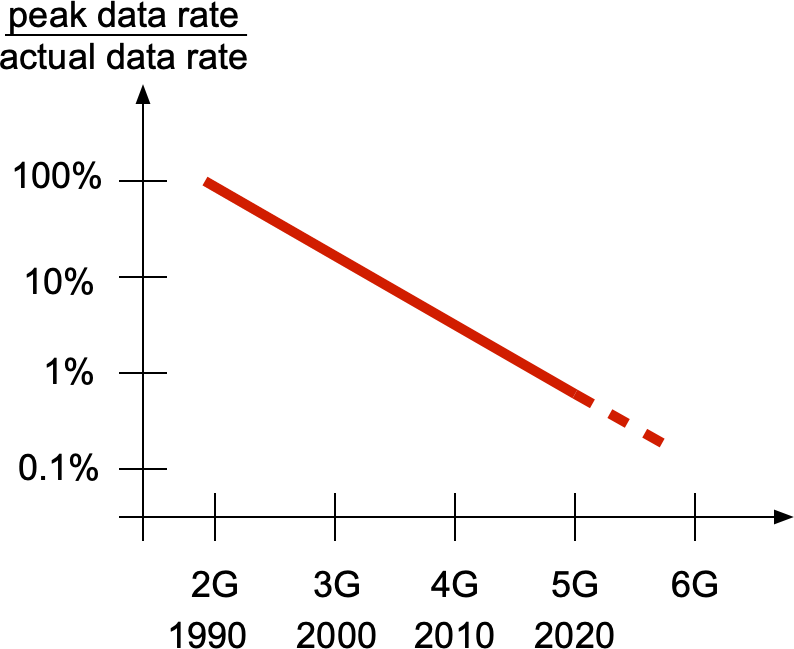
So even if 1 Tbps peak rates became a reality, users would likely experience between 1 Gbps to at most 10 Gbps – assuming their location is covered, which may vary over short spatial scales. Such user percentile performance can be analyzed using meta distributions.
In conclusion, while setting ambitious goals may trigger technological advances, it is important to be realistic of what is achievable and what performance the user actually experiences. For example, instead of focusing on 1 Tbps peak rates, we could focus on delivering 1 Gbps to 95% of the users, which may still be very challenging but probably achievable and more rewarding to the user. And speaking of billions of “connected devices” is just marketing unless it is clearly defined what being connected means.
For more information on the two analytical results above, please see this paper (Corollary 1) and this paper (Theorem 3).